RECTE Demonstrations with TRAPPIST-1 Observations¶
This Jupyter notebook demonstrates how to use RECTE to remove ramp effect systematics in the HST/WFC3 light curves. We apply RECTE correction to the TRAPPIST-1 observations (firstly published in De Wit et al. 2016). Results in this demosntration were published in Zhang et al. (2018).
Light curve Preparations¶
The light curve for each wavelength channel needs to be extracted from the ima frame before applying the RECTE correction (or any other systematics correction). The reference for this procedures is Deming et al. (2013). Light curves were pre-calculated for this demonstration and stored in a python shelve file.
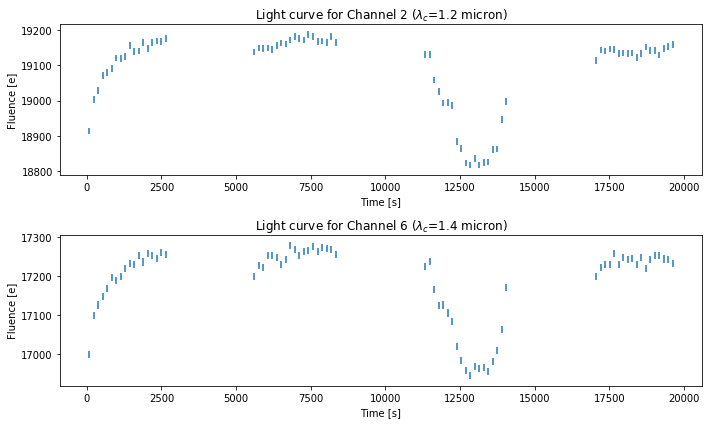
Uncorrected light curves¶
import shelve
import matplotlib.pyplot as plt
%matplotlib notebook
# First, restore light curve array from the shelve file
DBFileName = './demonstration_data/binned_lightcurves_visit_01.shelve'
saveDB = shelve.open(DBFileName)
LCarray = saveDB['LCmat']
ERRarray = saveDB['Errmat']
time = saveDB['time']
wavelength = saveDB['wavelength']
orbit = saveDB['orbit']
orbit_transit = np.array([2]) # transit occurs in the third orbits
expTime = saveDB['expTime']
saveDB.close()
# plot light curve of the second channel and the sixth channel
fig1 = plt.figure(figsize=(10, 6))
ax1 = fig1.add_subplot(211)
ax1.errorbar(time, LCarray[1, :], yerr=ERRarray[1, :], ls='none')
ax1.set_title('Light curve for Channel 2 ($\lambda_c$={0:.2} micron)'.format(wavelength[1]))
ax1.set_xlabel('Time [s]')
ax1.set_ylabel('Fluence [e]')
ax2 = fig1.add_subplot(212)
ax2.errorbar(time, LCarray[5, :], yerr=ERRarray[5, :], ls='none')
ax2.set_title('Light curve for Channel 6 ($\lambda_c$={0:.2} micron)'.format(wavelength[6]))
ax2.set_xlabel('Time [s]')
ax2.set_ylabel('Fluence [e]')
fig1.tight_layout()

RECTE corrections¶
We will use the convenience functions from the RECTECorrector module
to make corrections. Two functions RECTECorrrector1 and
RECTECorrector2 are for corrections for single-directional scanning
or “round-trip” scanning observations, respectively. The observation
used in this demonstration was done using “round-trip” scanning mode.
Therefore, we use RECTECorrector2 function to make the correction.
RECTECorrector2 function fits RECTE model profiles to the observed
light curves. It uses lmfit to perform the optimization. It excludes
the orbits where transits/eclipses occur. But this function can be
easily combined with transit model (such as batman) to include
orbits of transits/eclipses in fitting calculations. lmfit can also
be changed by MCMC sampler (such as emcee) to return more
informative fitting results.
In the following, we write a wrapper function removeRamp to get the
systematics-removed light curves
from RECTE import RECTE
from lmfit import Parameters, Model
from RECTECorrector import RECTECorrector2
def removeRamp(p0,
time,
LCArray,
ErrArray,
orbits,
orbits_transit,
expTime,
scanDirect):
"""
remove Ramp systemetics with RECTE
:param p0: initial parameters
:param time: time stamp of each exposure
:param LCArray: numpy array that stores all light curves
:param ErrArray: light curve uncertainties
:param orbits: orbit number for each exposure
:param orbits_transit: orbit number that transits occur. These orbits
are excluded in the fit
:param expTime: exposure time
:param scanDirect: scanning direction for each exposure. 0 for forward,
1 for backward
"""
nLC = LCArray.shape[0] # number of light curves
correctedArray = LCArray.copy()
correctedErrArray = ErrArray.copy()
modelArray = LCArray.copy()
crateArray = LCArray.copy()
slopeArray = LCArray.copy()
p = p0.copy()
for i in range(nLC):
correctTerm, crate, bestfit, slope = RECTECorrector2(
time,
orbits,
orbits_transit,
LCArray[i, :],
p,
expTime,
scanDirect)
# corrected light curve/error are normalized to the baseline
correctedArray[i, :] = LCArray[i, :] / correctTerm / (crate)
correctedErrArray[i, :] = ErrArray[i, :] / correctTerm / (crate)
modelArray[i, :] = bestfit
crateArray[i, :] = crate
slopeArray[i, :] = slope
return correctedArray, correctedErrArray, modelArray, crateArray, slopeArray
import pandas as pd
import numpy as np
infoFN = './demonstration_data/TRAPPIST_Info.csv'
info = pd.read_csv(infoFN)
grismInfo = info[info['Filter'] == 'G141']
scanDirect = grismInfo['ScanDirection'].values
p = Parameters()
p.add('trap_pop_s', value=0, min=0, max=200, vary=True)
p.add('trap_pop_f', value=0, min=0, max=100, vary=True)
p.add('dTrap_f', value=0, min=0, max=200, vary=True)
p.add('dTrap_s', value=50, min=0, max=100, vary=True)
LCarray_noRamp, ERRarray_noRamp, Modelarray, cratearray, slopearray = removeRamp(
p,
time,
LCarray,
ERRarray,
orbit,
orbit_transit,
expTime,
scanDirect)
Now, ramp systemetics are removed from the observations.
Result plot¶
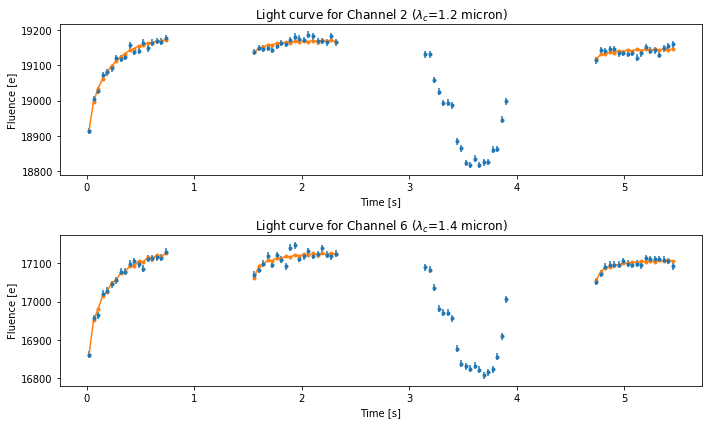
Best-fit Models¶
fig2 = plt.figure(figsize=(10, 6))
ax1 = fig2.add_subplot(211)
ax1.errorbar(
time / 3600,
LCarray[1, :],
yerr=ERRarray[1, :],
fmt='.',
ls='')
for o in [0, 1, 3]:
ax1.plot(
time[orbit == o] / 3600,
Modelarray[1, orbit == o],
'.-',
color='C1')
ax1.set_title('Light curve for Channel 2 ($\lambda_c$={0:.2} micron)'.format(wavelength[1]))
ax1.set_xlabel('Time [s]')
ax1.set_ylabel('Fluence [e]')
ax2 = fig2.add_subplot(212)
ax2.errorbar(
time / 3600,
LCarray[6, :],
yerr=ERRarray[6, :],
fmt='.',
ls='')
for o in [0, 1, 3]:
ax2.plot(
time[orbit == o] / 3600,
Modelarray[6, orbit == o],
'.-',
color='C1')
ax2.set_title('Light curve for Channel 6 ($\lambda_c$={0:.2} micron)'.format(wavelength[6]))
ax2.set_xlabel('Time [s]')
ax2.set_ylabel('Fluence [e]')
fig2.tight_layout()
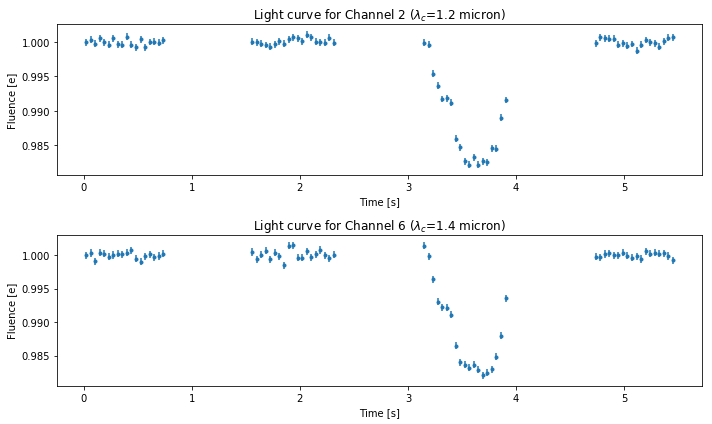
Corrected light curves¶
fig3 = plt.figure(figsize=(10, 6))
ax1 = fig3.add_subplot(211)
ax1.errorbar(
time / 3600,
LCarray_noRamp[1, :],
yerr=ERRarray_noRamp[1, :],
fmt='.',
ls='')
ax1.set_title('Light curve for Channel 2 ($\lambda_c$={0:.2} micron)'.format(wavelength[1]))
ax1.set_xlabel('Time [s]')
ax1.set_ylabel('Fluence [e]')
ax2 = fig3.add_subplot(212)
ax2.errorbar(
time / 3600,
LCarray_noRamp[6, :],
yerr=ERRarray_noRamp[6, :],
fmt='.',
ls='')
ax2.set_title('Light curve for Channel 6 ($\lambda_c$={0:.2} micron)'.format(wavelength[6]))
ax2.set_xlabel('Time [s]')
ax2.set_ylabel('Fluence [e]')
fig3.tight_layout()
